An example in HMM is a DNA sequence of nucleotides ACGT, where sequences can be modelled as the probabilities based on underlying hidden states, here Low GC content or High GC content. In this sequence analysis, it can apparantly be used to predict coding vs non-coding DNA regions. The example considers that H states actually characterize coding DNA, while L states characterize non-coding DNA states. How useful this actually is, or whether this is just a teaching example without any real applications, I am not sure! But it is an interesting example. We assume here that the probabilities stay constant. In the Markov assumption we assume that the next state depends only on the current state, and not any states before the current state.

AI generated picture of labelled DNA sequence. The labelling seems a bit off.
The transition matrix, here denoted by p, is given below. The emission matrix gives us the probabilities for each state, to observe each of the 4 nucleotides GCAT, with different probabilities for each. We have 2 states, and the matrix gives the probability to switch from H state to L state and the other way around.
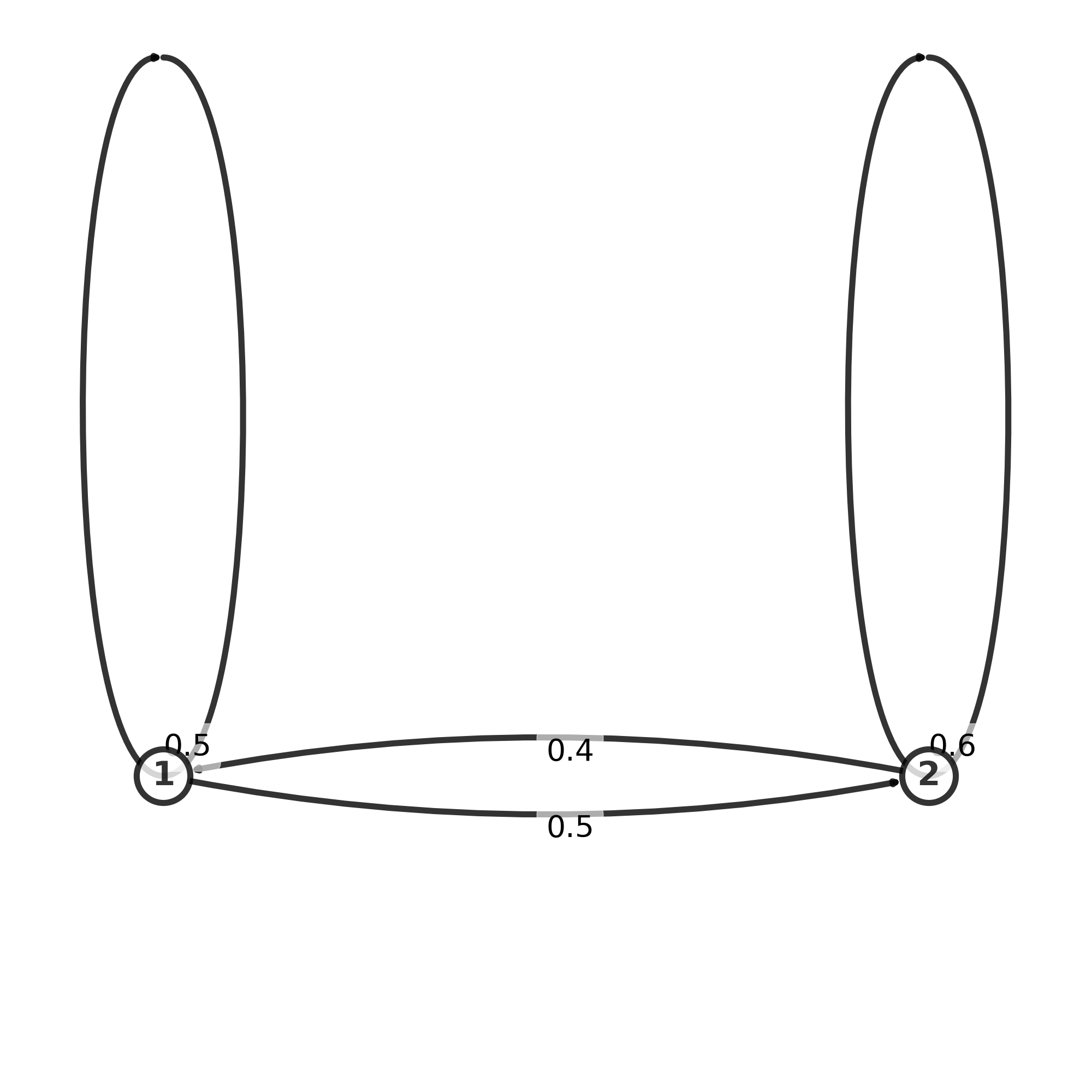
Graph of transmission matrix
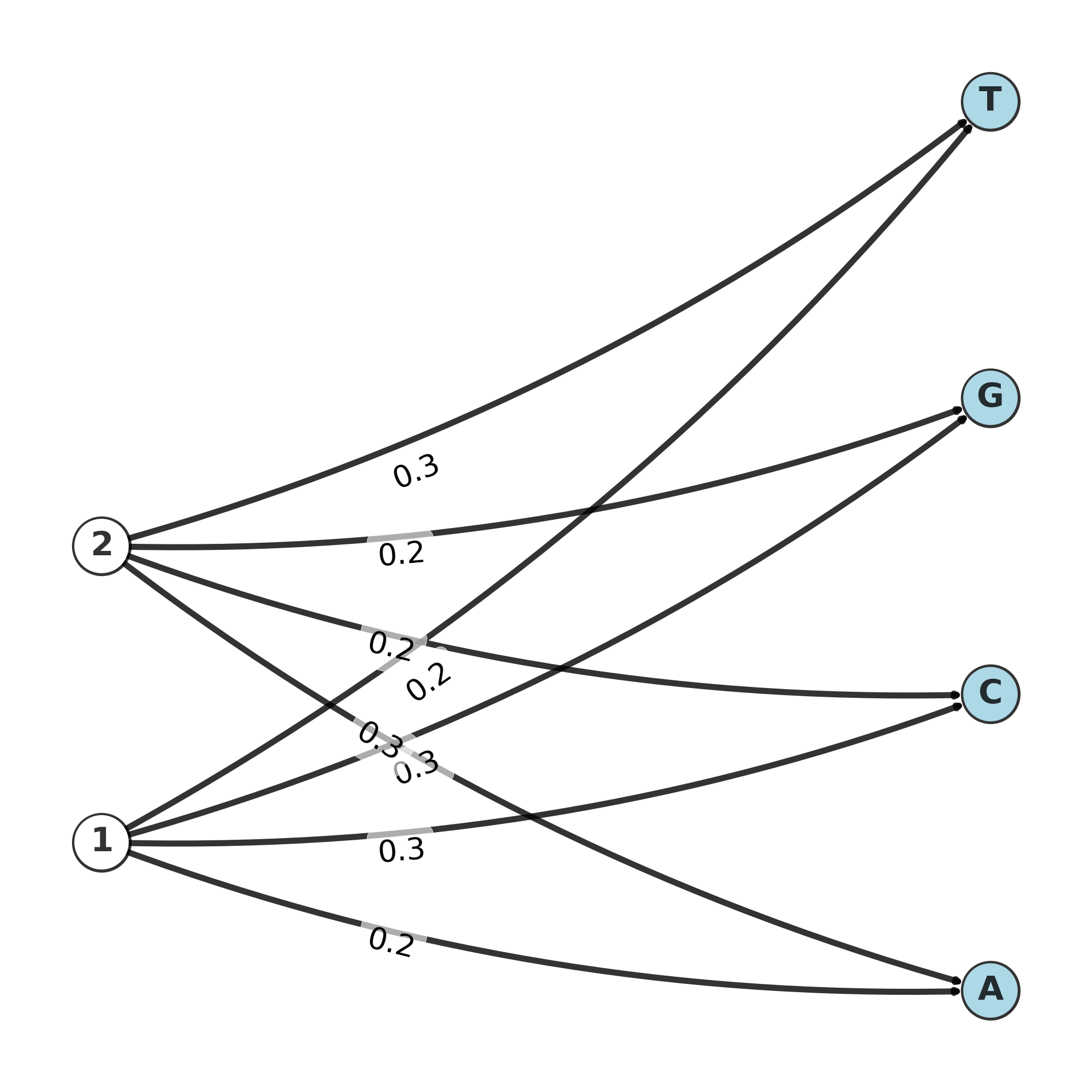
Graph of emission matrix

Transition matrix

Emission matrix
The nucleotides in the DNA sequence is what we actually observe, and finding out which sequence of Low/High states they were most likely drawn from, is the problem we are trying to solve. This is one of the three fundamental problems in Hidden Markov Models. We will use the Viterbi algorithm, which is actually a dynamic programming algorithm, since it solves the problem by looking at each step separately.
Steady state and eigenvectors
The Markov matrix is a stochastic matrix, so the largest eigenvalue will be 1. This means, over long time, applying the matrix p on an arbritary state vector, the result will converge to the eigenvector corresponding to the largest eigenvalue. The eigenvalues are seen below
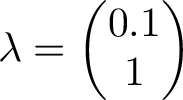
The eigenvector corresponding to the largest eigenvalue is the one to the right. We can compare this eigenvector with long term average of our simulation. The 2nd eigenvector will be be assumed positive components, when we compare with long-run simulation statistics.
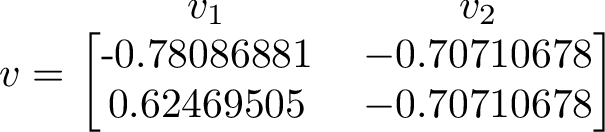
Counting how many times each state occurs and putting the counts in a vector let's us compare the right eigenvector to the long term simulated distribution. The counts from one run gave the values [0.70668522 0.70752809].
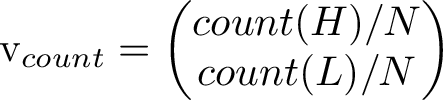
Scaling the long-term average vector to unit length and comparing it to the 2nd eigenvector, we see they are indeed very close to eachother!
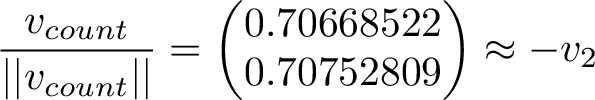
Obviously the count vector will have some random error from the simulation, and also the convergence to the 2nd eigenvector is only as the simulation goes on to infinity.
Viterbi algorithm - Most likely state sequence, from a given observation sequence
If we observe a sequence of emissions, and we know the p and e matrix, we can ask the question, which sequence of states was most likely to produce that observed sequence? Note, that many paths of states could give the same observed sequence, but one path would be the most probable. There could be several paths with the same maximum probability however, in that case, the Viterbi algorithm just outputs one such path, based on terminiation rules in the algorithm. The Viterbi algorithm was implemented based on the wikipedia page, and is a relatively short algorithm. Here it is implemented as written in wikipedia, however, variants include e.g taking the logarithm of probabilities, for numerical stability, especially for long sequences.
The probability of a path comes from multiplying together the elements of the transition and emission matrix for each timestep. In the end, a path with highest probability will be singled out by the algorithm. For a sequence of observations O and a sequence of states Q, the total joint probability of that outcome is given by
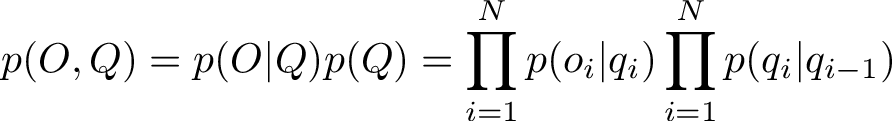
Note for the inital state i = 1 for q_1, we might select a uniform probability, in this case with 2 states we can set the initial probability as 1/2 for each. Alternatively, we could set the initial probability corresponding to the distribution in the eigenvector corresponding to the unit eigenvalue.
1) Observation test {GGCACTGAA}
Observing the sequence from the PDF example, we can find the highest probability path with Viterbi algorithm
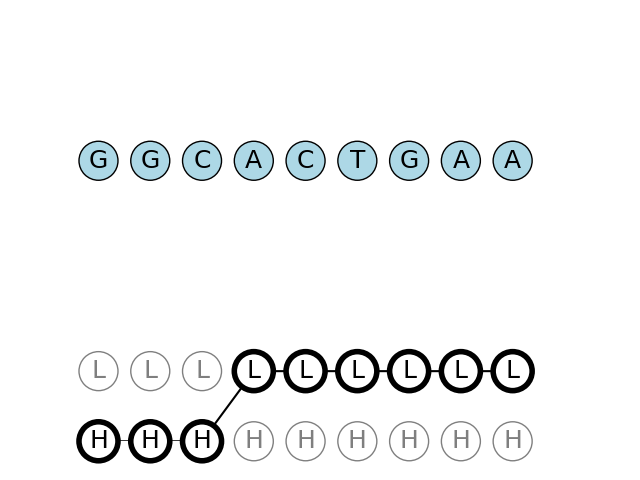
We can see that the path found is first the system was in H state, and then later, as T and more appears, for the system to have been in L state..
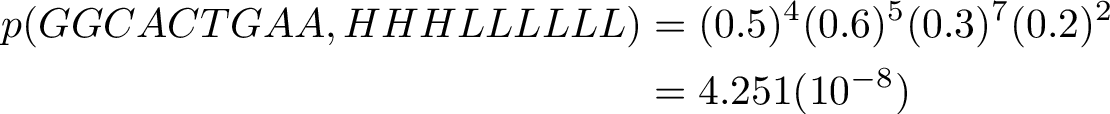
The probability of the states and observations is rather low, but it is the largest probability of other sequences that can give the same sequence of observations.
2) Observation test {AAAAA}
A total of 5 A nucleotides have been observed in the DNA string. We can find the path with the highest probability by using Viterbi algorithm
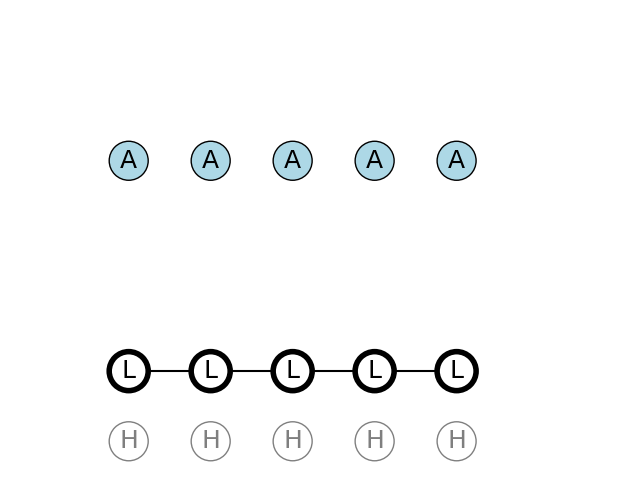
The max joint probability of sequence and observations is given below.
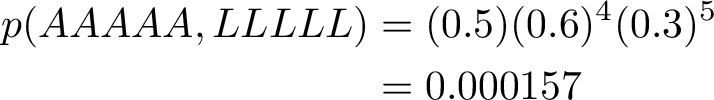
In this case the most probable path is picking only from the L state. We see that the probability to pick A nucleotide from the Low state is very high, and also, the probability to pick from the Low state on the next turn, after having picked from the Low state, is also very high.
3) Observation test {CCCCC}
We have observed 5 C nucleotides in a row. We use Viterbi algorithm to find the most probable path High/Low states
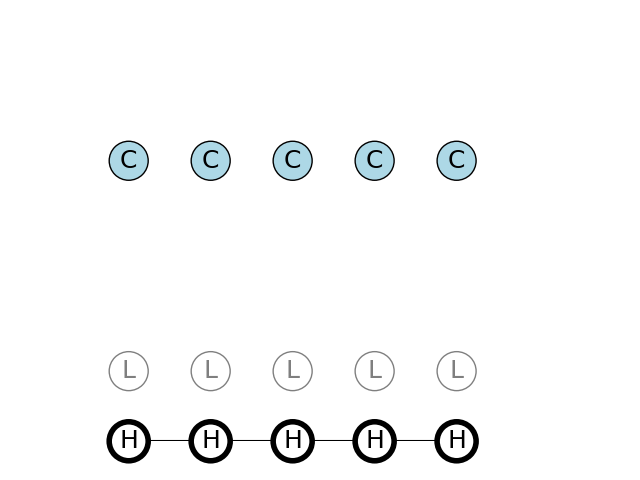
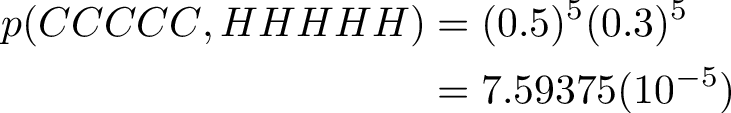
The most probable sequence of states to emit a sequence of C, will be a pure H state sequence. H stands for high G/C content. The calculated probability will be the same as for a sequence of 5 G nucleotides, as will be shown later.
4) Observation test {GGGGG}
We observe 5 G nucleotides in a row. We use Viterbi algorithm to find the most probable path of High/Low states
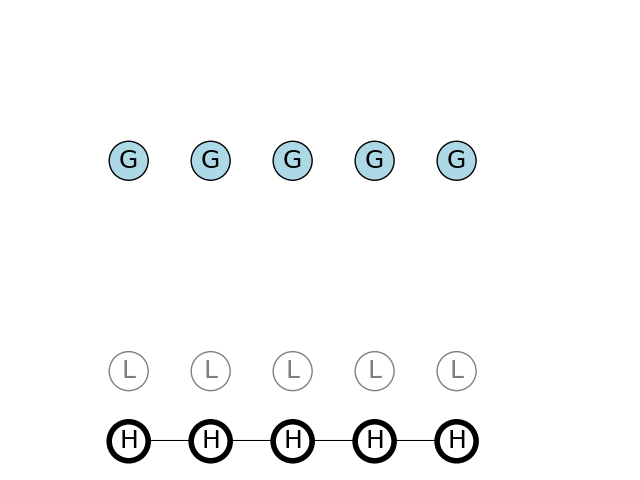
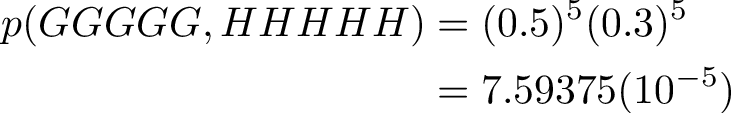
The path with highest probability for observing a sequence of G nucleotides, is a sequence of H states. They have a 50% chance to remain H on next step, and a high 30% chance of emitting G nucleotide.
5) Observation test {TTTTT}
We observe 5 T nucleotides in a row. We use Viterbi algorithm to find the most probable path of Low/High states.
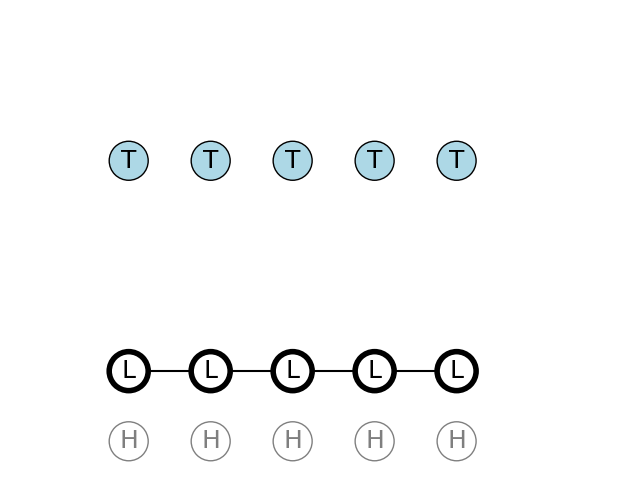
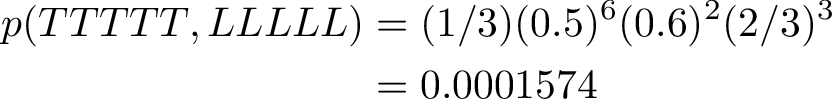
The path for only observing T is a sequence of L states. L state have a 60% chance to remain in L state on the next step, and also a high 30% chance of emitting a T nucleotide.
References
https://en.wikipedia.org/wiki/Viterbi_algorithm
https://www.cis.upenn.edu/~cis2620/notes/Example-Viterbi-DNA.pdf